For years, animal researchers have been able to rely on pure cell lines created by culturing different types of cells in the lab, a luxury not possible for plant researchers due to the plant cell wall. Now, for the first time, University of Georgia scientists have created a reference atlas that identifies multiple types of plant cells—and the DNA sequences that control them—at a molecular level, making it possible to study them without a cell line.
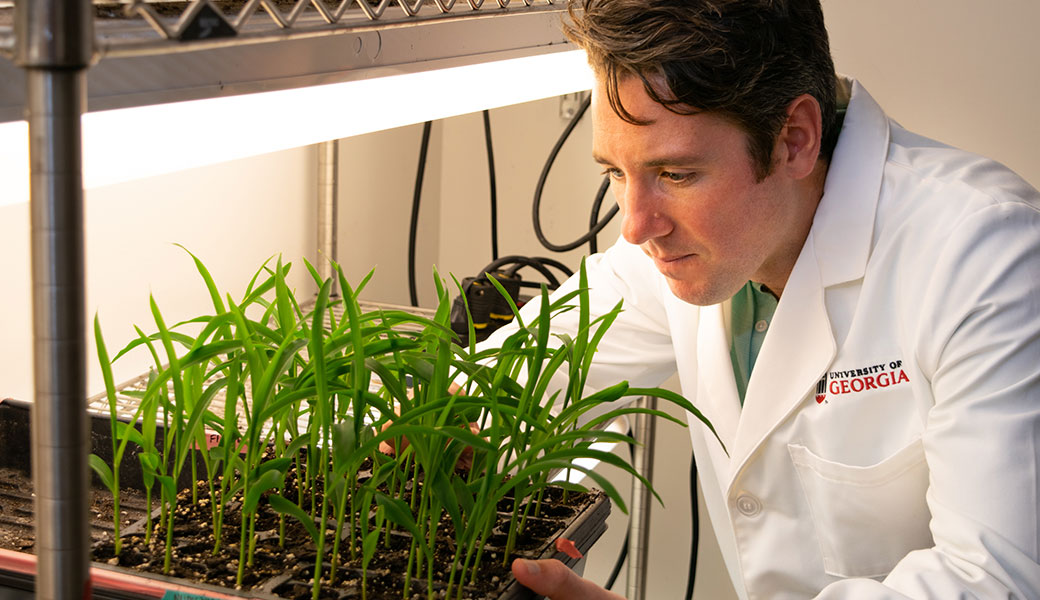
The project, led by Alexandre Marand, applied single-cell sequencing technology to break up tissues and organs from maize, isolating thousands of individual cells to determine the regulatory DNA—the parts of DNA that control the genes that are important in different types of cells.
“Plant cells are surrounded by a cell wall that is a significant hurdle for studying cell functions and development,” said Marand, first author and an NSF-funded postdoctoral researcher in the Schmitz lab. “Leveraging single-cell technology allowed us to bypass the technical challenges presented by the cell wall and determine the regulatory DNA signatures that define diverse types of cells at a scale that will be very useful for all plant scientists.”

Previously, experiments in plant science required complicated lab procedures taking years to generate transgenic lines to enable capture of very specific types of cells—one cell type at a time, according to the researchers. It required prior knowledge of genes with cell type-specific functions and was laborious, slow and expensive. The single-cell method samples cells unbiasedly, requires almost no labor and has a quick turnaround time with greater resolution of data.
“Alex’s research represents an important innovation considering almost nothing is known about how regulatory DNA controls cell identity and development in plants. With this technology, we can now learn about cellular development in any plant species,” said Bob Schmitz, associate professor of genetics in the Franklin College of Arts and Sciences. “You’re not limited to those that you’re able to make transgenics from.”
In the team’s study, published in Cell, the researchers focused on maize, creating a reference atlas of 52 cell types, more than 30 of which had never been described at a molecular level previously. Single-cell sequencing, which captures cells in between different states, allowed them to reconstruct their developmental history, delineating the molecular progression of one type of cell to another.

“While all cells in an organism generally have the same DNA, the parts that are active can change depending on the cell type,” Marand said. “Using statistical and computational methods, we’re able to reconstruct the developmental history of how maize organs are formed—which parts of the genome are important for coordinating the transition of one cell type into another. There are approximately 20 different types of cells in a leaf that all share a common origin. This technology allows us to separate those cells by their identity and retrace the molecular events that led to their current state.”
The researchers did this for 26 different types of developmental progressions in maize, as well as for Arabidopsis thaliana roots, a heavily studied model that allowed the team to validate their approach.
All of the team’s data is freely accessible via the National Center for Biotechnology Information, where users can access not just the raw data, but processed data that’s ready to be analyzed. The team also made available 52 maize cell type profiles, as well as those from Arabidopsis thaliana, on their genome browser.
“This is big data, so we’re talking about massive matrices that are thousands of gigabytes in size,” Schmitz said. “While anyone can download the data and reanalyze it, very few people are going to have the computational skills to be able to do that. But anyone is going to be able to go to our browser, look at their favorite gene and see which cell types regulatory DNA is active in.”
The results of the study confirmed that modern maize breeders have unwittingly been selecting for many of the regulatory sequences that the team identified in the paper. For Marand, the question is what’s next?
We think there are a lot of important regulatory DNA sequences with agronomic potential that haven’t been selected on yet,” he said. “This study gives a really nice list of places to explore to create more trait variation, particularly for genes we wouldn’t have considered before. The application of these data will have a big impact on biotechnology.”
Over the last decade, UGA has led the way in assembling plant genomes and determining the sequences important for each plant, according to Schmitz.
“Our lab depends on these genomes, as we specialize in figuring out how a plant interprets that sequence to respond to the environment or to produce a new cell type,” he said. “That’s the phase that the field is transitioning to right now. The phases are reading the genome, interpreting the genome, and I think the future is in writing the genome. Alex has made a landmark contribution to interpreting the genome by finding the parts that help control the genes in specific cell types.”
Researchers working at the bridge of biology and engineering are thinking about how to make bioproducts from plants, and to do that, they need lists of the genes and how those genes are controlled, Schmitz said.
“That’s where I think Alex’s work is going to be really important going forward,” he said. “Now that we are beginning to understand what the parts do, at least for maize, we’re in a position to begin writing the genome to fine-tune crop traits with big effects and to make them more valuable by bioengineering them to produce additional bioproducts. It’s not just maize—we’re also doing this for many other plants.”
Co-authors on the study include Zongliang Chen and Andrea Gallavotti, both at Rutgers University.
This study was funded with support from the NSF (IOS-1856627 and IOS-1802848) and the UGA Office of Research to Schmitz, the National Science Foundation (IOS-1916804) to Gallavotti, and an NSF Postdoctoral Fellowship in Biology (DBI-1905869) to Marand. Gallavotti and Schmitz are also supported by an NSF Collaborative award to support this research (IOS-2026554 and IOS-2026561).