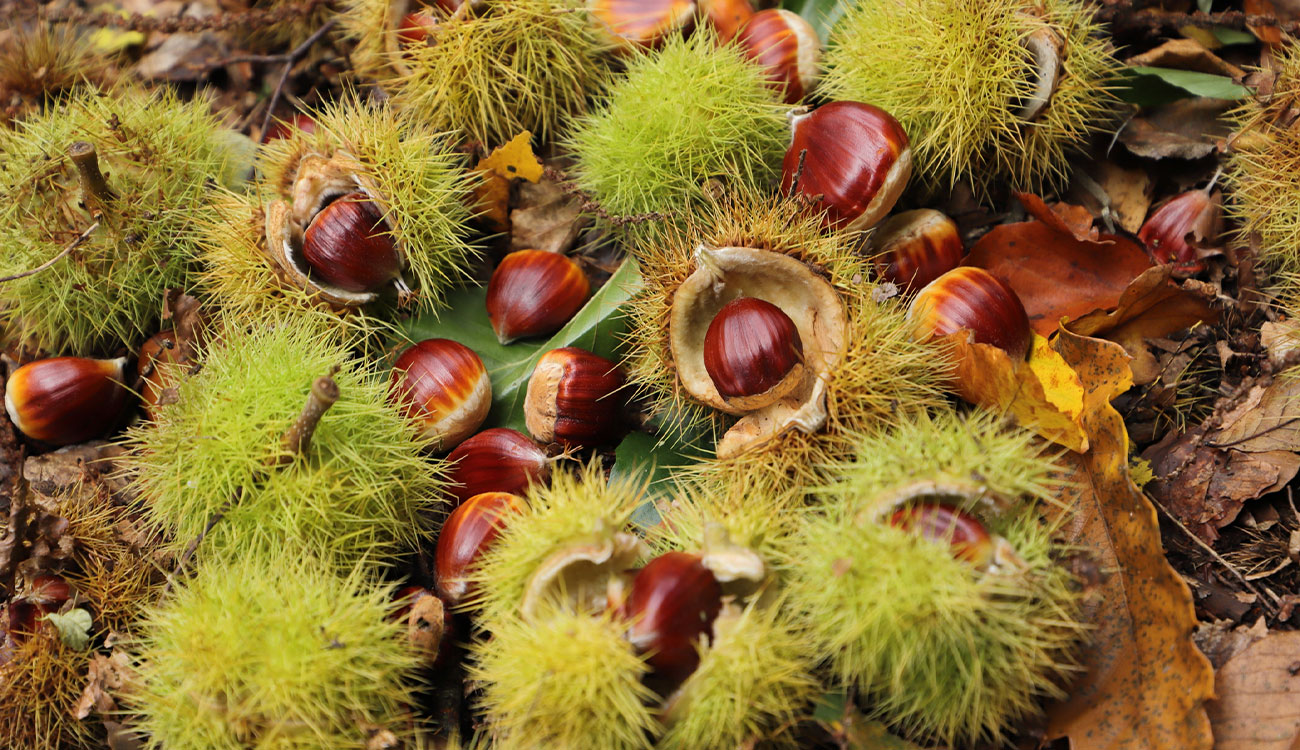
On a patch of land in Central New York last year, researchers collected more than 11,000 chestnuts.
But these nuts will absolutely not be roasted over an open fire. Instead, they will be replanted and tested for the presence of a specific gene, oxalate oxidase, or OxO, which gives the trees resistance to chestnut blight.
It’s still a long journey to replanting the billions of chestnuts lost over the past 100 years, when the fungus came to North America with an Asian chestnut tree. But it’s more progress than has been made in generations thanks to cellular-level technology, research partnerships and a technique for reproducing genetically identical chestnut embryos developed by scientists at the University of Georgia Warnell School of Forestry and Natural Resources.
Tough nut to crack
Chestnut trees, it turns out, are a finicky bunch.
When they began succumbing to blight in the early 1900s, scientists first tried attacking the pathogen, Cryphonectria parasitica. Scientists also tried to cross native chestnuts with Asian varieties, but these trees, even if they showed some resistance to blight, never persisted in the forest.

As technology evolved, scientists explored new ways of growing chestnuts. At Warnell in the 1980s, Professor Scott Merkle began investigating somatic embryogenesis to clone samples taken from surviving trees discovered in the wild. These trees lived long enough to produce nuts, which were harvested and used to start cultures. Around 1989 he fine-tuned the process: somatic embryogenesis starts with a seed embryo from a nut. It’s cloned to make thousands of new, genetically identical embryos, which proliferate until they are given a signal to complete their development into mature embryos. It’s these somatic embryos that germinate into seedlings.
At the time, scientists didn’t know what genes might be used to help chestnuts fight blight. Merkle thought the tissue cultures, if perfected on these finicky trees, could help down the road.
“He saw it as a tool for doing two things,” said Dayton Wilde, a professor in the UGA College of Agricultural and Environmental Sciences who worked in Merkle’s lab as a postdoctoral associate. “A target for genetic transformation—not even knowing what genes might be out there but knowing that there might be a resistance gene that could become available and there had to be a way to put it in—and second, once you get a tree with improved genetics you need to make lots of copies of it. You’re able to do mass propagation through embryogenesis.”
Around the same time, scientists at the State University of New York College of Environmental Sciences and Forestry (SUNY-ESF) were attempting to insert genes into new growth on shoots. It was as if the labs held pieces to the same puzzle. While Merkle worked to get somatic embryos to grow into seedlings as a vehicle for new genes, SUNY-ESF researchers Bill Powell and Chuck Maynard were experimenting with new genes, but couldn’t get them to take hold in the plants.
“At that time, we had a good genetic transformation system for working with yellow poplar embryogenic cultures and sort of adapted it for chestnut. Scott really was the first to develop a somatic embryogenesis system for chestnut,” said Wilde. “About that time, SUNY-ESF was also interested in chestnut restoration and wanted to collaborate with Scott on this.”
Maynard had recruited postdoctoral researcher Zizhuo Xing, who helped link the puzzle pieces together: He applied Merkle’s somatic protocol to work with embryogenic cultures.
The missing link
In a 2019 interview SUNY-ESF’s Powell gave for the Templeton World Charity Foundation, he described the connection he made between oxalic acid and chestnut blight—the fungus uses oxalic acid to attack the tree, but if you can grow a tree that breaks down oxalic acid, it, in theory, is no longer susceptible to blight.
Using a natural bacterial process to insert the OxO gene into cells grown in a lab, they were able to grow chestnuts with a level of resistance similar to Asian chestnuts.
“Once the bacteria move the DNA into plant cells, we propagate those cells and ultimately grow trees from those cells. Scott’s work has really been key to that process of getting from a single cell to a whole plant—especially for chestnut,” said Andy Newhouse, co-director of the American Chestnut Research and Restoration Project at SUNY-ESF. “Part of the reason this whole restoration effort for chestnuts with genetic engineering has taken more than 30 years is that every step of the process has to be considered. But Scott’s work and overlapping work in our labs has allowed us to make it happen.”
Over the decades, the friendly competition between the labs helped drive new discoveries. Seedling company ArborGen and, more recently, the Forest Health Initiative also provided funding for both labs to work on somatic embryogenesis and gene transfer systems.

The American Chestnut Foundation has also been a key partner in this work. It’s helped fund research, formalize processes and create a network of nurseries willing to help in the restoration process.
The advances just in the past couple of decades have been astounding, said Sara Fitzsimmons, chief conservation officer for The American Chestnut Foundation.
“Up until the mid-2010s there were lots of people working on various aspects of chestnut restoration, and there was collaboration through a group called NE-1333—we would get together and see where there was cross-collaboration,” said Fitzsimmons. “Scott’s been focusing primarily on tissue culture and cryopreservation, figuring out the mechanisms of improving the transgenic pipeline. Bill’s lab at ESF has been doing that with a lot of input on what’s been going on at Scott’s lab.”
Long-awaited approval
There’s still one more roadblock before the chestnuts in Central New York can be found at a nursery near you, though. And that has to do with genes and regulations.
When Powell’s lab successfully transformed a New York American Chestnut with the OxO gene, it created a new line called Darling 58. They then produced somatic embryos from this line, rooting the shoots to produce transgenic Darling 58 trees.
When transgenic trees are used in research, they require a field test permit from USDA-APHIS. While this is a complex process, it’s not nearly as involved as the “non-regulated status” required to distribute a transgenic tree to the public. This requires approval from the USDA-APHIS, the EPA and the FDA.
It’s been years in the making, but a decision is expected in the next few months.
If approved, it’s a major step for transgenic trees. Until now, the process has been used by the agricultural sector for transgenic crop approvals. Companies hire teams to complete and shepherd the applications.
“We were told it wasn’t feasible,” added Newhouse, who has seen the federal application through to its final stages. “And it did take years of meetings and figuring things out, but here we are. We have engagement from all three agencies, and we expect a decision soon.”
Merkle sighs when he recalls the paperwork required to grow transgenic chestnut trees at Whitehall Forest or even to pollinate chestnuts at the UGA Horticultural Farm a couple years ago. He used pollen from SUNY-ESF; the trees produced nuts, which were then sent to SUNY-ESF, Penn State University and the TACF research farm in Meadowview to be tested for blight resistance.
As for his own crop of transgenic trees, Merkle said he’s done with field testing. But he’s perfectly happy to work in his lab growing transgenic seedlings for others to field test. And that’s exactly how Merkle is helping with the next stage of chestnut development.
Improving the herd
While chestnuts grew from Maine to Georgia, the varying climates and locations created different genotypes across the range—slightly different genetics in response to local conditions. Because future chestnuts are selected for the OxO gene, it means a block of genes adjacent to the OxO gene on the chromosome of Darling 58 is likely to get carried along with each generation, rather than being diluted over time. “So, there’s some unknown-size chunk of that New York tree that’s going to be produced in all the OxO trees we produce here in Georgia using Darling 58 pollen,” said Merkle.

He is now working to directly insert the OxO gene into regional varieties of chestnuts, which can then be grown in their native climates if Darling 58 receives federal approval. This work, supported by the American Chestnut Foundation and the Foundation for the Carolinas, is being done in Merkle’s lab by research professional Ryan Tull and postdoctoral associate Heather Gladfelter. Research professional Paul Montello oversees cryostoring copies of the cultures for future use.
This work will continue to strengthen the future of American chestnuts, said Jared Westbrook, director of science for the American Chestnut Foundation.
“Scott’s been working with us for the past few years on putting the OxO gene in other trees, not just trees from New York, so we don’t have a bottleneck (of genes),” said Westbrook. “Scott is the pioneer for developing these methods for cultivating chestnut embryos and doing some of the genetic engineering—cloning chestnuts in a way that we could put in different genes into the tree’s genome.”
The goal, he said, is to breed the OxO gene into 200 trees from across the range. Each year scientists hand-pollinate a few thousand trees. With deregulation, they could expand to planting 10,000 trees a year.
Rough estimates put the original population of chestnuts at 4 billion. That’s a lot of trees to replace, and a large-scale logistical problem for Fitzsimmons, who is charged with getting the trees back into the forest.
But working with nurseries, in anticipation of potentially distributing seedlings later this year, has her hopeful. It’s a sea change from where chestnuts were just a couple decades ago.
“Chestnuts are what we call a recalcitrant species—the seed is only storable for a year, and there’s all these things that make it a difficult species to work with,” she said. “But that’s some of the leaps and bounds Scott’s and Bill’s labs have made. They’ve found ways to overcome some of these challenges.”
This story was first published by the Warnell School of Forestry & Natural Resources and is republished with permission.