Antibiotic resistance is an increasing health problem, but new research suggests it is not only caused by the overuse of antibiotics. It’s also caused by pollution.
Using a process known as genomic analysis, University of Georgia scientists found a strong correlation between antibiotic resistance and heavy metal contamination in an environment.
Jesse C. Thomas IV, an alumnus of the College of Public Health and the Savannah River Ecology Laboratory, found commonalities in soils contaminated with heavy metals on the U.S. Department of Energy’s Savannah River Site near Aiken, South Carolina.
According to the study, recently published in the journal Microbial Biotechnology, soils with heavy metals had a higher level of specific bacterial hosts that were accompanied by antibiotic-resistant genes.
Antibiotic-resistant genes
Hosts included Acidobacteriaoceae, Bradyrhizobium and Streptomyces. The bacteria had antibiotic-resistant genes, known as ARGs, for vancomycin, bacitracin and polymyxin. All three drugs are used to treat infections in humans.
The bacteria also had an ARG for multidrug resistance, a strong defense gene that can resist heavy metals as well as antibiotics, according to Thomas, who was conducting his doctoral research at the time.
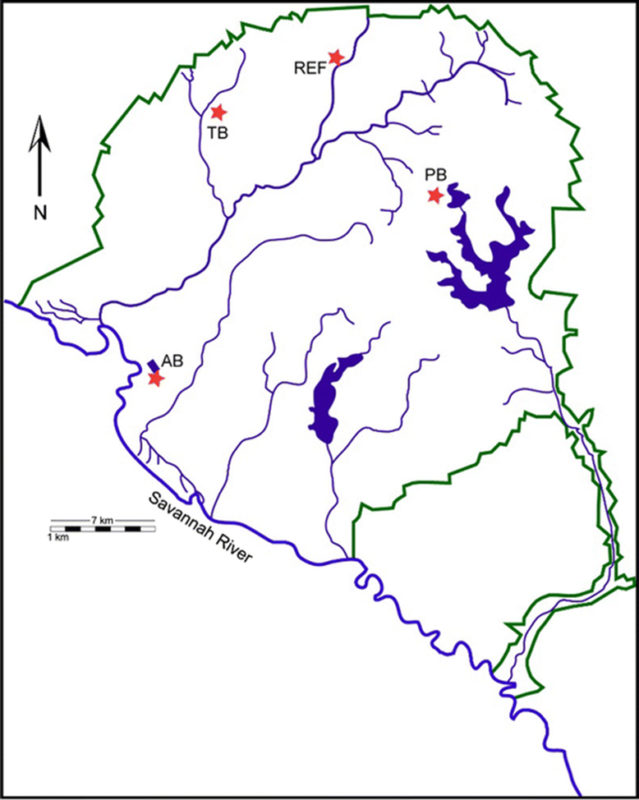
When these ARGs were present in the soil, metal-resistant genes, or MRGs, were present for several metals including arsenic, copper, cadmium and zinc.
Thomas, currently a biologist at the Centers for Disease Control and Prevention, said microorganisms develop new strategies and countermeasures over time to protect themselves.
“The overuse of antibiotics in the environment adds additional selection pressure on microorganisms that accelerates their ability to resist multiple classes of antibiotics. But antibiotics aren’t the only source of selection pressure,” Thomas said. “Many bacteria possess genes that simultaneously work on multiple compounds that would be toxic to the cell, and this includes metals.”
Exerting long-standing pressure
Travis Glenn, professor in the public health college, advised Thomas during the study. He said more research needs to be done to determine if metal-resistant genes respond in the same way to bacteria as antibiotic-resistant genes.
Unlike antibiotics, heavy metals don’t degrade in the environment so “they can exert long-standing pressure,” according to Glenn, who also directs the Institute of Bioinformatics.
The study reports previous research identified antibiotic-resistance in heavy metal-contaminated streams on the site by examining water samples in the lab.
“When you expose the sample to a drug on a petri dish or assay, it only represents a fraction. This doesn’t give you a complete picture. With genomic analysis we were able to get much further,” Thomas said.
Impact on water and food
The significance of the research is they can start to characterize bacterial communities and specific ARG and MRG genes in the environment, Glenn said.
It is clear that there are several human pathogens that develop antibiotic resistance — overuse is not the only cause, according to Thomas. Human activities like agriculture and the combustion of fossil fuels play a role.
“We need a better understanding of how bacteria are evolving over time,” he said. “This can impact our drinking water and our food and eventually our health.”
Additional authors on the study include Adelumola Oladeinde, Bacterial Epidemiology and Antimicrobial Resistance Research Unit, USDA, Athens, Georgia; Troy J. Keiran, University of Georgia, College of Public Health, John W. Finger Jr., Auburn University, Department of Biological Sciences, Auburn, Alabama; Natalia J. Beyona-Vasquez, UGA College of Public Health and the Institute of Bioinformatics; John C. Cartee, Division of STD Prevention, CDC, Atlanta; James C. Beasley, UGA Savannah River Ecology Laboratory, Aiken, South Carolina, Warnell School of Forestry and Natural Resources, Athens, Georgia; Olin E. Rhodes Jr., SREL and Odum School of Ecology, Athens, Georgia; John C. Seaman and J. Vaun McArthur, SREL.